RNA G-Quadruplex (rG4) – What it is, why it matters, how to study it
What is RNA G-Quadruplex (rG4)?
RNA G-quadruplexes (rG4s) are non-canonical secondary structures originated from guanine-rich RNA sequences, where four guanines are arranged together by Hoogsteen hydrogen bonding in a planar G-quartet and the quartets are stacked forming a stable four-stranded conformation. Monovalent cations, particularly K⁺, greatly enhance the rG4 stability (Fig. 1). rG4s are evolutionarily conserved and are prevalent in the human transcriptome, with a significant enrichment in regulatory regions such as 5′ and 3’ UTRs, introns, and non-coding RNAs (ncRNAs). Approximately 13,000 potential human rG4-forming sequences have been computationally predicted. However, their actual folding into rG4s in vivo is dynamic and context-dependent, being modulated by RNA-binding proteins (RBPs), cations, and cellular stressors.
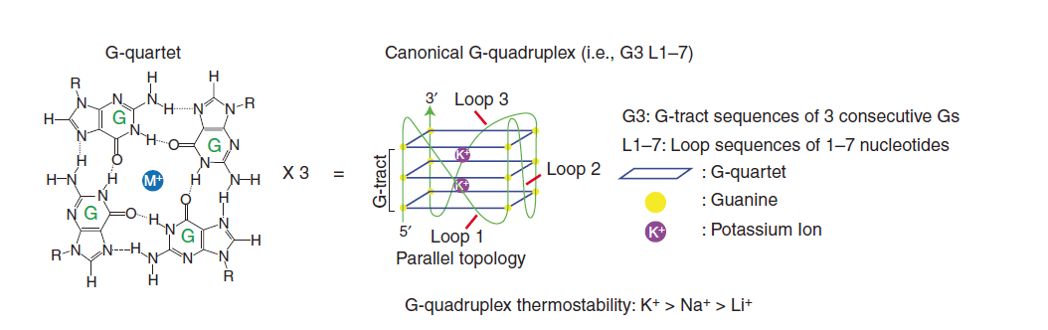
Figure 1. RNA G-quadruplex (rG4) structure. The chemical structure of G-quartet, showing Hoogsteen hydrogen bonds (dotted lines) between guanines (G) and monovalent cation (M+, e.g. K+) (Left). Three tiered G-quartets are stacked to form a canonical rG4, with connecting G-track and loop sequences between the G-quartet planes (Right)[1].
Why Are rG4s Important in Biology and Disease?
rG4s regulate gene expression at multiple levels of mRNA translation regulation, mRNA stability, pre-mRNA splicing and transcription, RNA phase separation and localization. rG4s are also closely linked to diseases, such as cancer, neurodegeneration, and stress response. These roles make rG4s both functional regulators and potential therapeutic targets.
- rG4 regulation of mRNA translation
Dynamic rG4 folding/unfolding have the potentials to regulate mRNA translation (Fig. 2-7)[2, 3], often suppressing cap-dependent translation, but also enabling IRES-mediated translation.
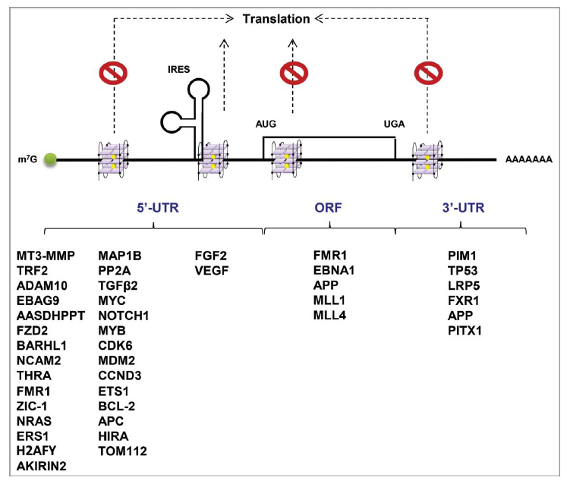
Figure 2. rG4s in mRNA translation regulation. rG4s in the 5′-UTRs, ORFs, and 3′-UTRs predominantly inhibit cap-dependent translation. However, rG4s near internal ribosome entry site (IRES) in 5′-UTR are likely to augment IRES-mediated translation. Target mRNA examples known to be regulated by rG4s are listed underneath [3].
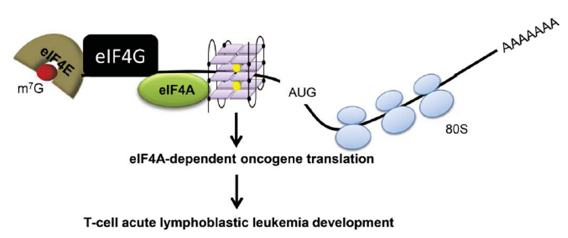
Figure 3. rG4s in 5’UTR regulate eIF4A-dependent oncogene translation. eIF4A unwinds rG4 in the 5’-UTR of many key transcription factor and oncogene mRNAs for translation, thereby contributing to the T cell-acute lymphoblastic leukemia development [4].
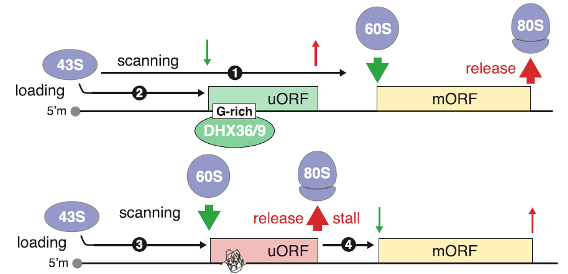
Figure 4. rG4s in 5’UTR select uORF/mORF for translation. When 43S preinitiation complex (PIC) scans through unfolded rG4 sequence and unstructured 5’-UTR (e.g. rG4 unfolded by DHX36/9 helicases), the translation initiates at the main open reading frame (mORF). However, the PIC scanning can be slow down by rG4 structure in the 5’UTR (e.g. in the absence of rG4 helicases), increasing the likelihood of upstream open reading frame (uORF) translation. uORF translation causes 80S ribosomes stall and dissociation, preventing further downstream mORF translation [5].
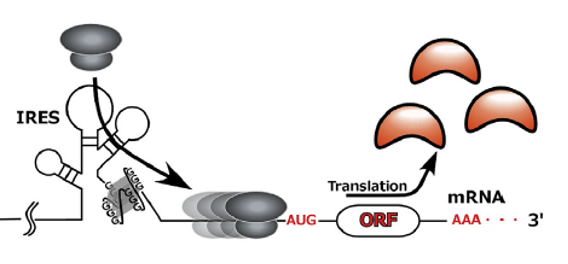
Figure 5. rG4s in 5’UTR act as internal ribosome entry site (IRES). rG4s in FGF2, α-Syn and VEGF mRNAs are proposed to stimulate translation as a part of an internal ribosome entry site (IRES) or an IRES-like element, by helping recruit the 40S ribosome [6].
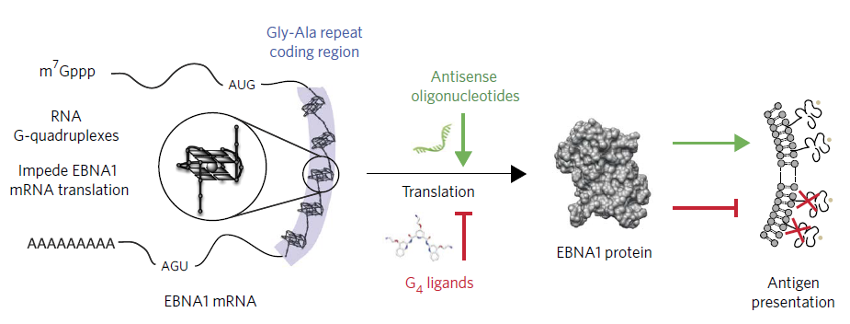
Figure 6. rG4s in ORF regulate translation and immune invasion. EBNA1, an Epstein–Barr virus encoded protein, is finely tuned for its expression via rG4 in its ORF to down regulate protein translation: high enough for sustaining viral infection in the cells but not too high as presented viral antigen to illicit host immune response. Antisense oligonucleotides target and disrupt the rG4s, boosting EBNA1 synthesis, viral antigen presentation, and host immune response. Conversely, a small-molecule ligand (PDS) stabilizes the rG4s, reducing EBNA1 mRNA translation and antigen presentation [7].
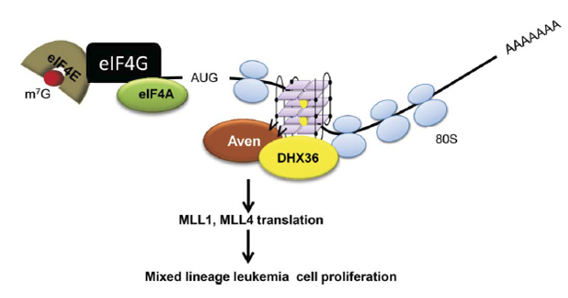
Figure 7. rG4s in ORF stimulate translation. In the ORF of MLL1 or MLL4 mRNA, the rG4 can bind methylated Aven protein to recruit DHX36 helicase. DHX36 unwinds the rG4 and facilitates polysome translation. Ultimately, the rG4 promotes the translation of oncogenic MLL1 and MLL4 proteins, thereby enhancing leukemic cell proliferation [8].
- rG4 regulation of mRNA stability
rG4s affect miRNA binding and degradation of mRNA, controlling the RNA stability(Fig. 8)[4].
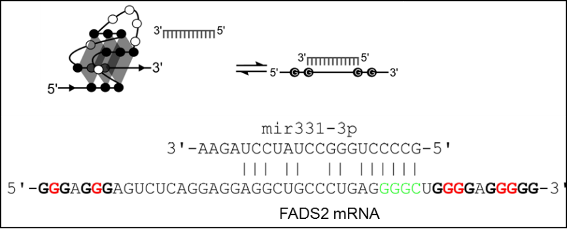
Figure8. rG4s in 3’UTR regulate miRNA binding and mRNA stability. Folding of rG4 in miRNA recognition element (MRE) in the 3’UTR of mRNA can prevent miRNA targeting and silencing of the mRNA, as in this case of the equilibrium between rG4 in FADS2 mRNA and mir331-3p binding in cellulo [9].
- RNA phase separation and localization
Especially in neuronal cells, rG4s can act as a signal for mRNA subcellular targeting in neurite [10] and form scaffolds for droplet phase separation in the cells (Fig. 9) [11]. In addition, rG4 in the pre-mRNA can affect pre-mRNA maturation, including 3’-end processing for poly-A tail addition and intron removal through splicing (Fig. 10)[12].
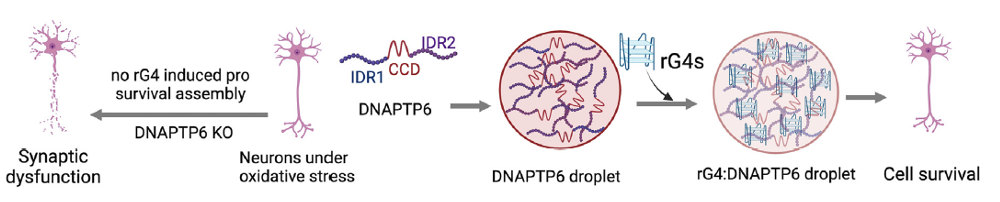
Figure 9. rG4s in mRNA form scaffolds for phase separation. Under stress, rG4-bearing mRNAs interact with DNAPTP6 to promote stress granule (SG) assembly in neurons. These mRNAs are recruited to SGs in an rG4-dependent manner through their interaction with DNAPTP6, which induces RNA self-assembly, phase separation, and cell survival [11].
- RNA splicing and transcription
rG4 structures in pre-mRNAs influence the RNA processing and chromatin interactions, thereby regulating splicing and transcription (Fig.10)[12].
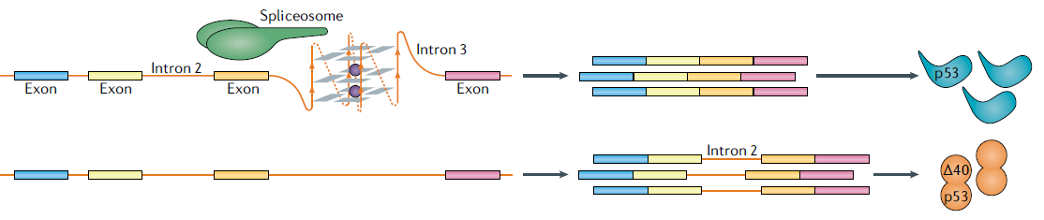
Figure 10. rG4s in pre-mRNA affect splicing. The interaction between rG4 and spliceosome-associated RNA-binding proteins orchestrates the splicing process of adjacent introns, exemplified by the second intron of the p53 gene [12].
Dynamic rG4 folding/unfolding in lncRNAs can regulate chromatin modifier function (Fig. 11)[13], RNA transcription (Fig. 12)[14], decoying functional molecules (e.g. proteins) (Fig. 13)[15], and pre-miRNA processing(Fig. 14)[16] in the nucleus.
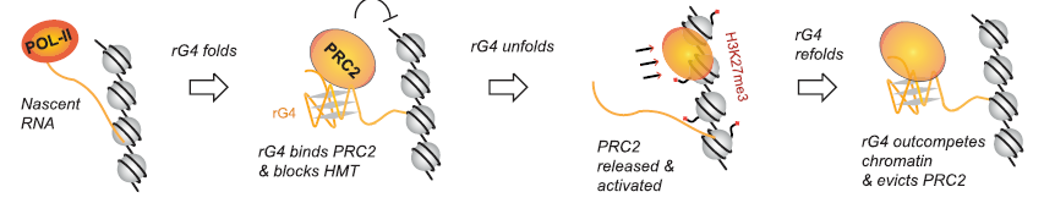
Figure 11. Dynamic folding and unfolding of rG4 regulate PRC2 recruitment, activation, and eviction. The formation of rG4 on nascent transcripts of Xist lncRNA attracts PRC2 chromatin modifier, inhibiting the histone methyltransferase activity. As the RNA competes with DNA for PRC2 binding, PRC2 is prevented from loading onto chromatin. When rG4 unfolds, PRC2 is released from the RNA to load onto chromatin, where its histone methyltransferase activity is activated for H3K27me3 modification. This process is reversible, with the refolding of rG4 draws PRC2 back to the RNA still in close spatial proximity, thereby evicting PRC2 from the chromatin [13].
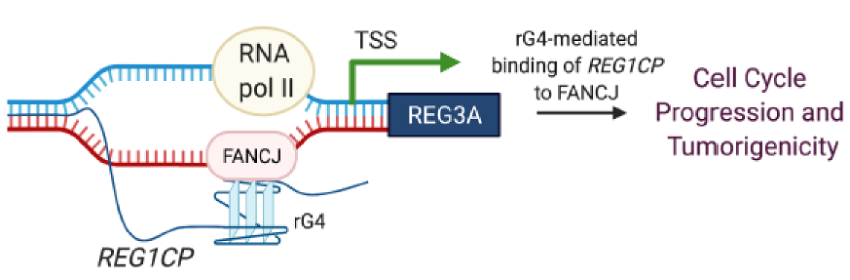
Figure 12. rG4 folding regulates gene transcription. The binding of rG4 in REG1CP lncRNA to FANCJ helicase unwinds double-stranded DNA in the TSS, relieves transcriptional inhibition, and turns on REG3A gene transcription [14, 17]. The expressed REG3A promotes cancer cell cycle progression and tumorigenicity.
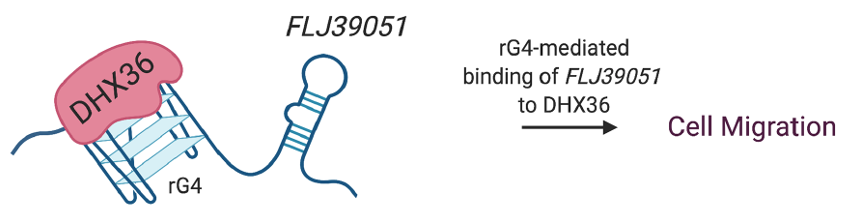
Figure 13. rG4s in lncRNA decoy RNA helicase DHX36. The rG4 in FLJ39051 lncRNA interacts with DHX36 RNA helicase and inhibits the ATP-dependent rG4 unwinding activity of DHX36, thereby promoting the motility of colorectal cancer cells [15].
- rG4 interplay with epitranscriptomic modifications
rG4s are known to interplay with RNA epitranscriptomic modifications, such as m7G (Fig. 14)[16], o8G (Fig. 16)[18], and m6A (Fig. 15)[19, 20], to co-regulate gene expression.
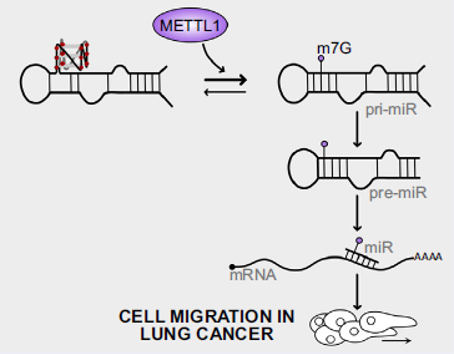
Figure 14. rG4s and its epitranscriptomic modification regulate pri-miRNA processing. The rG4 in pre-miRNA impedes the pre-miRNA transition to canonical conformation, thereby hindering the maturation of pre-miRNA. In contrast, m7G modification by METTL1 can counteract the rG4 structure, allowing processing to mature oncogenic miRNA and cancer cell migration [16].
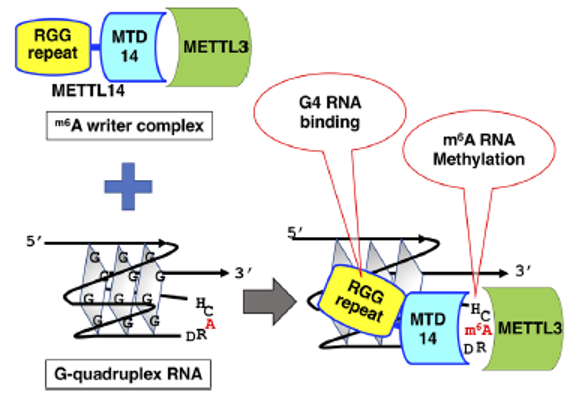
Figure 15. The m6A methyltransferase heterodimer, composed of METTL3 and METTL14, exhibits a binding preference for rG4 structures through the RGG repeats of METTL14. This heterodimer selectively methylates adenosines in close proximity to rG4 sequences. These findings imply a potential mechanism by which the METTL3/METTL14 complex is directly recruited to specific methylation sites, particularly those near G4-forming regions.
- rG4s in diseases
Dynamic rG4 folding/unfolding are involved in stress response to modulate cellular adaptation (Fig. 16)[18], cancer gene expression regulation (Fig. 17)[21, 22], and neurodegenerative diseases such as Parkinson’s disease via α-synuclein protein aggregation (Fig. 18), dementia with Lewy bodies, and multiple system atrophy[23].
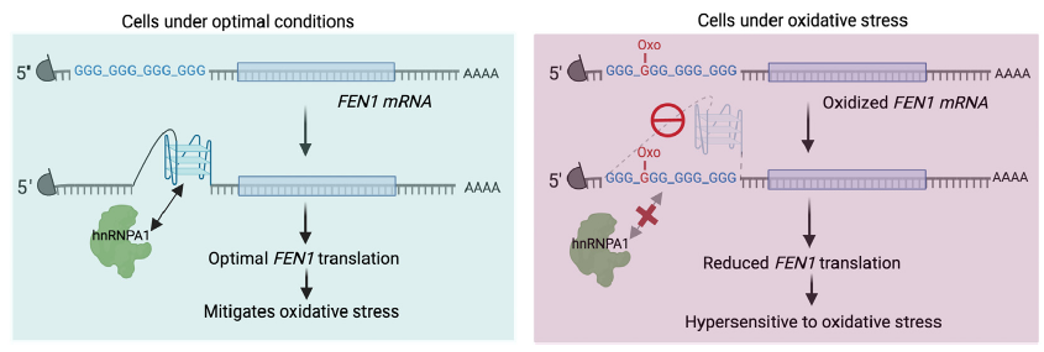
Figure 16. rG4 in oxidative stress. Oxidation of the guanine base (o8G) in rG4-forming sequence in oxidative stress-responsive FEN1 mRNA can impede the formation of rG4. This disruption can eliminate the interaction between heterogeneous nuclear ribonucleoprotein A1 (hnRNPA1) and rG4, thereby inhibiting the translation of FEN1. Consequently, the cells become more susceptible to oxidative stress [18, 17].
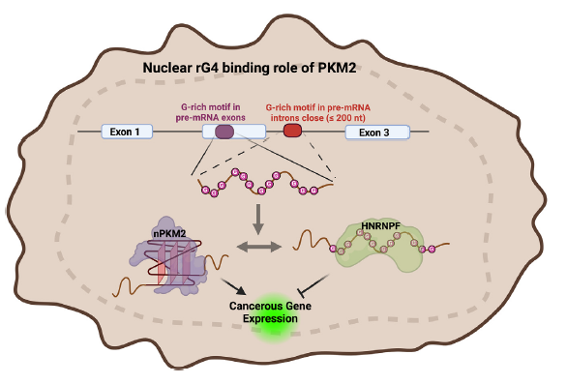
Figure 17. rG4 folding/unfolding in pre-mRNA controls transcriptional output of rG4-containing RNAs (rG4ome) in cancer. Guanine-rich regions in the exons and introns of many cancerous precursor mRNAs can be either unfolded in a linear conformation or folded into rG4 structures. In cancer cells, the competition between HNRNPF, which recognizes and stabilizes the linear form of G-rich motifs, and nuclear PKM2, which recognizes and stabilizes the G-quadruplex form of G-rich motifs, regulates the expression of cancer-related genes [21, 22].
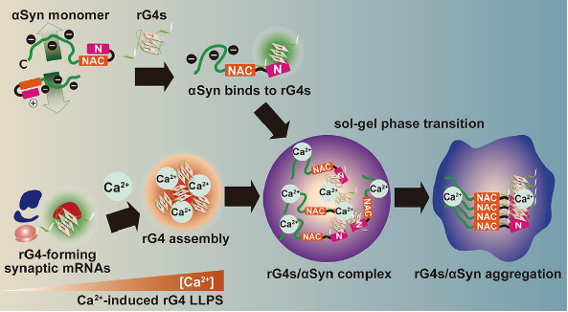
Figure 18. rG4s as scaffolds for α-synuclein aggregation. rG4 assembly, inducible by calcium influx, can accelerate the phase transition and serve as scaffolds for neuropathological aggregation of α-synuclein [23]. Synucleinopathies, such as Parkinson’s disease, dementia with Lewy bodies, and multiple system atrophy, are initiated by the aggregation of α-synuclein and lead to progressive neurodegeneration.
How to Profile rG4s in Your Research
While rG4s are increasingly recognized as key regulatory elements, their detection remains technically challenging. Arraystar rG4 Microarray is a powerful tool to profile in vivo rG4 landscape and dynamics[24]. It uses in vivo DMS treatment to capture, rG4 immunoprecipitation to enrich, and microarray to sensitively detect/quantify rG4s, much better than RNA-seq especially for rG4 RNAs often at low abundance.

References
[1] Kwok, C.K., et al. (2018) “Detecting RNA G-Quadruplexes (rG4s) in the Transcriptome” Cold Spring Harb Perspect Biol 10(7):[PMID:29967010]
[2] Arora, A. and Suess, B. (2011) “An RNA G-quadruplex in the 3′ UTR of the proto-oncogene PIM1 represses translation” RNA Biol 8(5):802-5 [PMID:21734463]
[3] Song, J., et al. (2016) “RNA G-quadruplexes and their potential regulatory roles in translation” Translation (Austin) 4(2):e1244031 [PMID:28090421]
[4] Wolfe, A.L., et al. (2014) “RNA G-quadruplexes cause eIF4A-dependent oncogene translation in cancer” Nature 513(7516):65-70 [PMID:25079319]
[5] Murat, P., et al. (2018) “RNA G-quadruplexes at upstream open reading frames cause DHX36- and DHX9-dependent translation of human mRNAs” Genome Biol 19(1):229 [PMID:30591072]
[6] Leppek, K., et al. (2018) “Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them” Nat Rev Mol Cell Biol 19(3):158-174 [PMID:29165424]
[7] Murat, P., et al. (2014) “G-quadruplexes regulate Epstein-Barr virus-encoded nuclear antigen 1 mRNA translation” Nat Chem Biol 10(5):358-64 [PMID:24633353]
[8] Thandapani, P., et al. (2015) “Aven recognition of RNA G-quadruplexes regulates translation of the mixed lineage leukemia protooncogenes” Elife 4([PMID:26267306]
[9] Rouleau, S., et al. (2017) “3′ UTR G-quadruplexes regulate miRNA binding” RNA 23(8):1172-1179 [PMID:28473452]
[10] Subramanian, M., et al. (2011) “G-quadruplex RNA structure as a signal for neurite mRNA targeting” EMBO Rep 12(7):697-704 [PMID:21566646]
[11] Asamitsu, S., et al. (2023) “RNA G-quadruplex organizes stress granule assembly through DNAPTP6 in neurons” Sci Adv 9(8):eade2035 [PMID:36827365]
[12] Varshney, D., et al. (2020) “The regulation and functions of DNA and RNA G-quadruplexes” Nat Rev Mol Cell Biol 21(8):459-474 [PMID:32313204]
[13] Lee, Y.W., et al. (2024) “G-quadruplex folding in Xist RNA antagonizes PRC2 activity for stepwise regulation of X chromosome inactivation” Mol Cell 84(10):1870-1885 e9 [PMID:38759625]
[14] Yari, H., et al. (2019) “LncRNA REG1CP promotes tumorigenesis through an enhancer complex to recruit FANCJ helicase for REG3A transcription” Nat Commun 10(1):5334 [PMID:31767869]
[15] Tassinari, M., et al. (2021) “Biological relevance and therapeutic potential of G-quadruplex structures in the human noncoding transcriptome” Nucleic Acids Res 49(7):3617-3633 [PMID:33721024]
[16] Pandolfini, L., et al. (2019) “METTL1 Promotes let-7 MicroRNA Processing via m7G Methylation” Mol Cell 74(6):1278-1290 e9 [PMID:31031083]
[17] Kharel, P. and Ivanov, P. (2024) “RNA G-quadruplexes and stress: emerging mechanisms and functions” Trends Cell Biol 34(9):771-784 [PMID:38341346]
[18] Ma, Y., et al. (2023) “RNA G-Quadruplex within the 5′-UTR of FEN1 Regulates mRNA Stability under Oxidative Stress” Antioxidants (Basel) 12(2):[PMID:36829835]
[19] Yoshida, A., et al. (2022) “Recognition of G-quadruplex RNA by a crucial RNA methyltransferase component, METTL14” Nucleic Acids Res 50(1):449-457 [PMID:34908152]
[20] Jara-Espejo, M., et al. (2020) “Potential G-Quadruplex Forming Sequences and N(6)-Methyladenosine Colocalize at Human Pre-mRNA Intron Splice Sites” ACS Chem Biol 15(6):1292-1300 [PMID:32396327]
[21] Anastasakis, D.G., et al. (2024) “Nuclear PKM2 binds pre-mRNA at folded G-quadruplexes and reveals their gene regulatory role” Mol Cell 84(19):3775-3789 e6 [PMID:39153475]
[22] Kharel, P. and Ivanov, P. (2024) “PKM2-G-quadruplex interactions conspire to regulate the cancer transcriptome” Mol Cell 84(19):3574-3575 [PMID:39366344]
[23] Matsuo, K., et al. (2024) “RNA G-quadruplexes form scaffolds that promote neuropathological alpha-synuclein aggregation” Cell 187(24):6835-6848 e20 [PMID:39426376]
[24] rG4 Array Service [https://arraystar.com/services/rg4-array-service/]
