Small RNA NGS Solutions
Related Products
Related Services
Related Reviews
m7G TRAC-Seq
TRAC-Seq (tRNA reduction and cleavage sequencing) is a chemically-based method for unbiased, single-nucleotide resolution profiling of m7G modifications throughout the tRNA transcriptome. Arraystar m7G TRAC-Seq service offers the sample-to-data solution. With the wealth of information from the m7G TRAC-seq profiling, investigators can easily advance their next step research in this field of study.
Benefits
• Single-nucleotide resolution: Precise mapping of m7G sites at nucleotide-level accuracy.
• Chemical specificity: Highly specific chemical reactions, instead of antibody affinity, to eliminate background from non-specific antibody binding and enable quantitative stoichiometry assessment.
• Superior tRNA coverage: Demethylation step using Arraystar rtStar™ tRNA Pretreatment Kit to overcome tRNA modification barriers that block reverse transcription, allowing efficient sequencing of heavily modified tRNAs
• Comprehensive analyses: Simultaneous identification of m7G sites, and discovery of modification-associated sequence motifs (e.g. RAGGU motif for m7G tRNAs)
| Service Name | Price |
|---|---|
| m7G TRAC-Seq Service |
N7-methylguanosine (m7G) is a conserved tRNA modification predominantly located at position 46 in the variable loop of certain tRNAs, enzymatically deposited by METTL1/WDR4 methyltransferase complex. It plays a pivotal role in maintaining tRNA structural integrity through formation of a C13–G22–m7G46 base triple, thereby shielding tRNAs from rapid decay pathways. Functionally, m7G is indispensable for stem cell self-renewal and proper lineage commitment during embryonic development [1] . Moreover, m7G-modified tRNAs enhance the translation efficiency of codon-enriched mRNAs, particularly those encoding oncoproteins involved in cell cycle progression, proliferation, and growth signaling. m7G dysregulation, especially with METTL1 overexpression, elevates levels of m7G-modified tRNAs (e.g. Arg-TCT-4-1, Lys-CTT, Val-AAC), driving tumorigenesis in diverse malignancies including leukemia, glioblastoma, cholangiocarcinoma, and lung cancer [2-4] .
References
1. Lin S et al: Mettl1/Wdr4-Mediated m(7)G tRNA Methylome Is Required for Normal mRNA Translation and Embryonic Stem Cell Self-Renewal and Differentiation. Mol Cell 2018, 71(2):244-255 e245.[PMID: 29983320]
2. Orellana EA et al: METTL1-mediated m(7)G modification of Arg-TCT tRNA drives oncogenic transformation. Mol Cell 2021, 81(16):3323-3338 e3314.[PMID: 34352207]
3. Dai Z et al: N(7)-Methylguanosine tRNA modification enhances oncogenic mRNA translation and promotes intrahepatic cholangiocarcinoma progression. Mol Cell 2021, 81(16):3339-3355 e3338.[PMID: 34352206]
4. Ma J et al: METTL1/WDR4-mediated m(7)G tRNA modifications and m(7)G codon usage promote mRNA translation and lung cancer progression. Mol Ther 2021, 29(12):3422-3435.[PMID: 34371184]
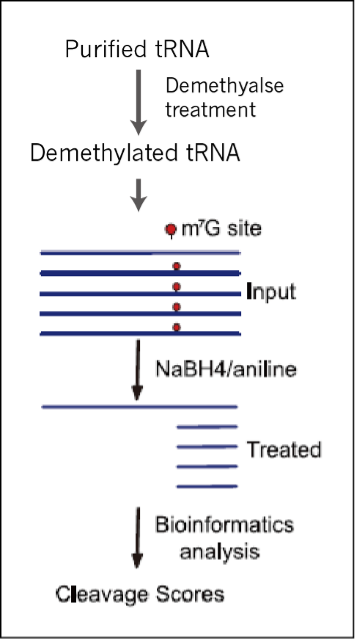
Figure 1. m7G TRAC-Seq workflow. Isolated tRNAs are enzymatically demethylated and treated with NaBH 4 /aniline to induce reductive cleavage specifically at m7G residues. The 3’ cleavage fragments are ligated with a sequencing adaptor at their 5’-ends where the precise modification sites are located. The generated cDNA library for each sample is sequenced. The sequencing data are analyzed for m7G sites and modification levels by cleavage scores.
The m7G TRAC-seq includes detailed bioinformatics analyses to facilitate insights into m7G in tRNA biology, diseases, and biomarker applications.
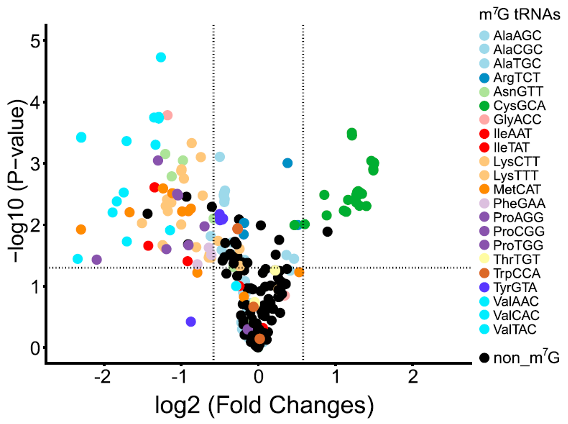
Volcano plot for differentially expressed m7G-tRNAs
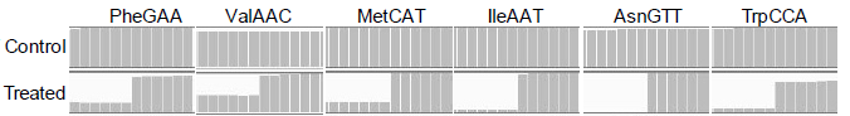
m7G TRAC-seq read alignments for m7G site identification.
Motif analysis of m7G site identified by TRAC-seq.

